Japanese
次元削減
次元削減は高次元データから潜在的な特徴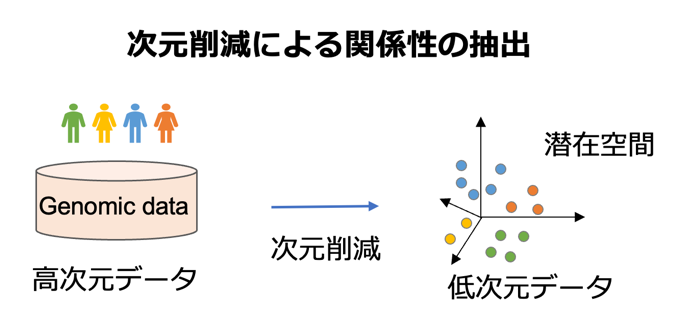 や性質を保存した低次元空間を得る技術です。 次元削減法として主成分分析などが有名ですが、本研究室では高次元空間上での局所的な関係を保存するLaplacian Eigenmapなどをはじめとしたアルゴリズムとその高度化の研究を行っています。
や性質を保存した低次元空間を得る技術です。 次元削減法として主成分分析などが有名ですが、本研究室では高次元空間上での局所的な関係を保存するLaplacian Eigenmapなどをはじめとしたアルゴリズムとその高度化の研究を行っています。
クラスタリング
クラスタリングは教師ラベルを与える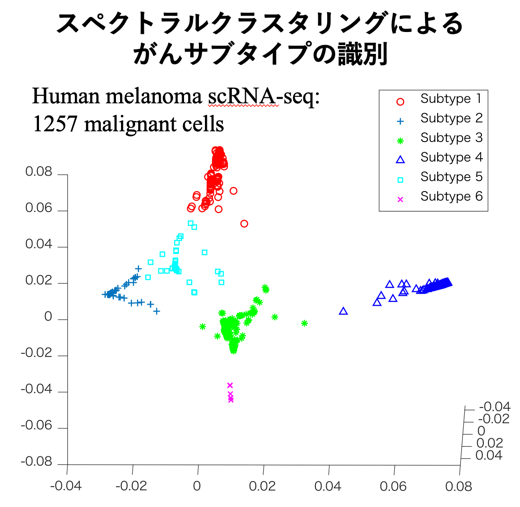 ことなく、データを似ている者同士があつまるグループ(クラスタ)に分けるタスクです。クラスタリングには様々なアルゴリズムがありますが、スペクトラルクラスタリングなどの上記の次元削減手法と関係が深いアルゴリズムの研究を行っています。また医療・生命分野やマテリアルズインフォマティクスの分野との連携した応用技術開発も進めています。
ことなく、データを似ている者同士があつまるグループ(クラスタ)に分けるタスクです。クラスタリングには様々なアルゴリズムがありますが、スペクトラルクラスタリングなどの上記の次元削減手法と関係が深いアルゴリズムの研究を行っています。また医療・生命分野やマテリアルズインフォマティクスの分野との連携した応用技術開発も進めています。
深層学習技術の高度化
深層学習ではどのような学習手法を使うかでその性能を左右しますが、本研究室では通常用いられる確率的勾配降下法系の手法ではなく、非線形非負値行列分解を使った学習手法を開発しています。さらに同手法と確率的勾配降下法とのハイブリッド型手法なども開発しています。またそれらの高性能計算環境向けの実装の高度化にも取り組んでいます。
生命・医療データへの応用
生命・医療分野の研究者・医師・企業等と連携して生命・医療データへの各種機械学習応用技術の開発を行っています。遺伝子発現データ、代謝産物データ、ゲノム変異データといった様々なデータを扱い、開発したアルゴリズムや理論の実応用に取り組んでいます。
English
Dimensionality reduction
Dimensionality reduction is a technology to transf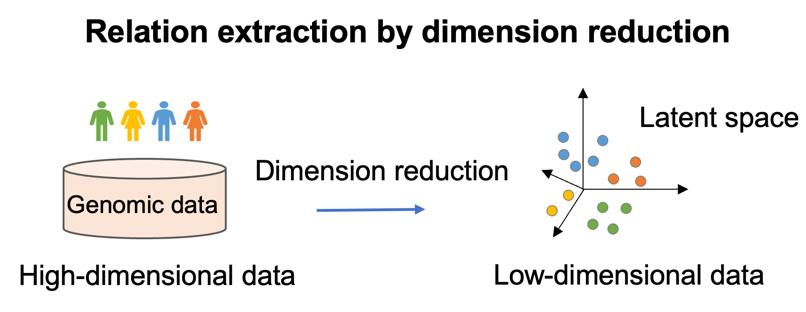 orm data from a high-dimensional space into a low-dimensional space so that the low-dimensional representation retains some meaningful and latent properties of the original data. In our Lab, we focus on dimensionality reduction methods that preserve the local relationships in high-dimensional space, such as locality preserving projection (LPP) and Laplacian Eigenmap-based methods.
orm data from a high-dimensional space into a low-dimensional space so that the low-dimensional representation retains some meaningful and latent properties of the original data. In our Lab, we focus on dimensionality reduction methods that preserve the local relationships in high-dimensional space, such as locality preserving projection (LPP) and Laplacian Eigenmap-based methods.
Clustering
Clustering is an important technique for exploratory 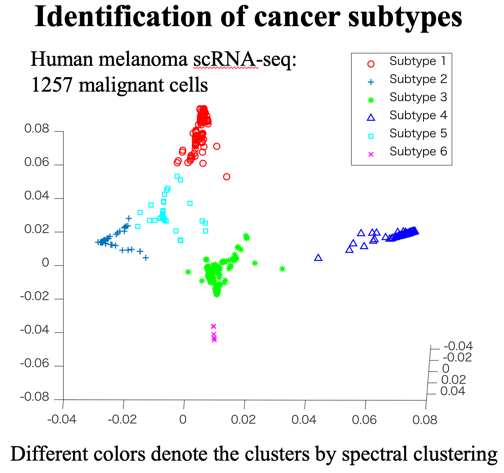 data analysis with the objective of grouping unlabeled data objects in the same cluster which are similar to each other. We are focusing on the algorithms that are closely related to the above dimensionality reduction methods such as spectral clustering. We are also developing clustering technologies in collaboration with the medical, life, and materials informatics fields.
data analysis with the objective of grouping unlabeled data objects in the same cluster which are similar to each other. We are focusing on the algorithms that are closely related to the above dimensionality reduction methods such as spectral clustering. We are also developing clustering technologies in collaboration with the medical, life, and materials informatics fields.
Advanced deep learning technology
In deep learning, the performance depends on what kind of learning method is used. In our laboratory, instead of the usually used stochastic gradient descent method, we have developed a learning method that uses non-negative matrix factorization. We are also developing a hybrid method of the non-negative matrix factorization and the stochastic gradient descent. We are also working on a high-performance implementation for parallel computing environments.
Application to life science and medical data
We are developing various machine learning application technologies for life science and medical data in collaboration with researchers, medical doctors, companies, etc. We handle various data such as gene expression data, metabolite data, and genomic variation data, and are working on the practical application of the developed algorithms and theories.
